Generating Configurations
Instead of writing PostgreSQL schemas, Elasticsearch mappings, and Arranger configs by hand, the Stage portal includes a Config Generator that produces all configuration files from your CSV data in one step.
Navigate to Config Generator in the Stage portal navigation bar (visible once Stage is running).
Step 1: Provide CSV Data
Upload a .csv file using the Upload .csv file button, or paste CSV content directly into the text area. Once loaded, a preview of the first five rows is shown so you can confirm the correct file was used.
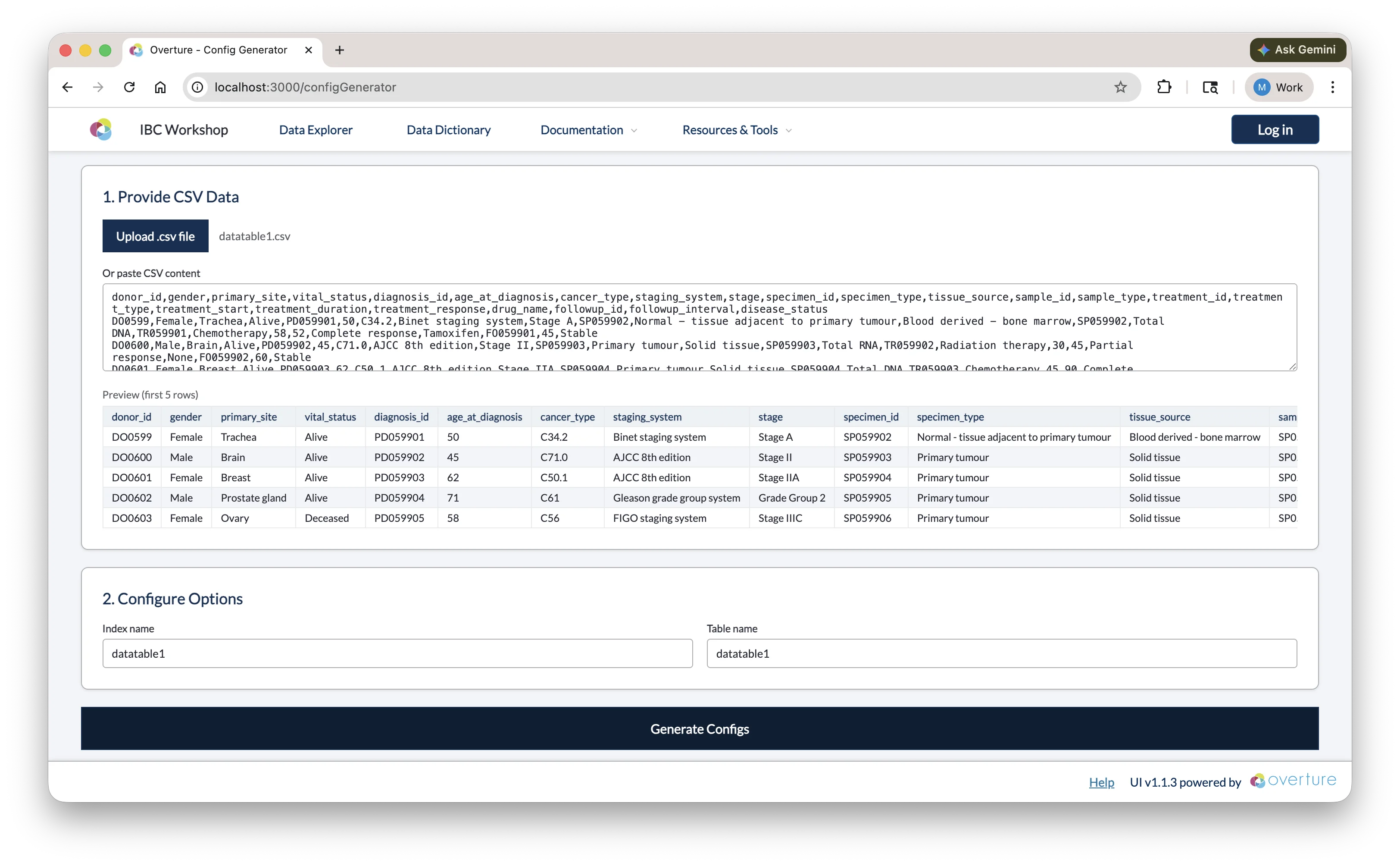
Step 2: Configure Options
| Field | Description |
|---|---|
| Index name | The name of the Elasticsearch index. Auto-populated from the CSV filename; edit if needed (e.g. datatable1). |
| Table name | The name of the PostgreSQL table. Defaults to the same value as the index name. |
Step 3: Generate and Copy
Click Generate Configs. Once complete, the output panel shows a tabbed view with seven files:
| Tab | File | Where to save |
|---|---|---|
postgres-table.sql | CREATE TABLE statement | setup/configs/postgresConfigs/ |
elasticsearch-mapping.json | Index template and field mappings | setup/configs/elasticsearchConfigs/ |
arranger/base.json | Arranger index and document type | setup/configs/arrangerConfigs/<indexName>/ |
arranger/extended.json | Human-readable field display names | setup/configs/arrangerConfigs/<indexName>/ |
arranger/table.json | Data table column configuration | setup/configs/arrangerConfigs/<indexName>/ |
arranger/facets.json | Sidebar filter configuration | setup/configs/arrangerConfigs/<indexName>/ |
lectern/dictionary.json | Lectern data dictionary | setup/configs/lecternDictionary/ |
Use the Copy button on each tab to copy the content, then paste it into the corresponding file in your project.
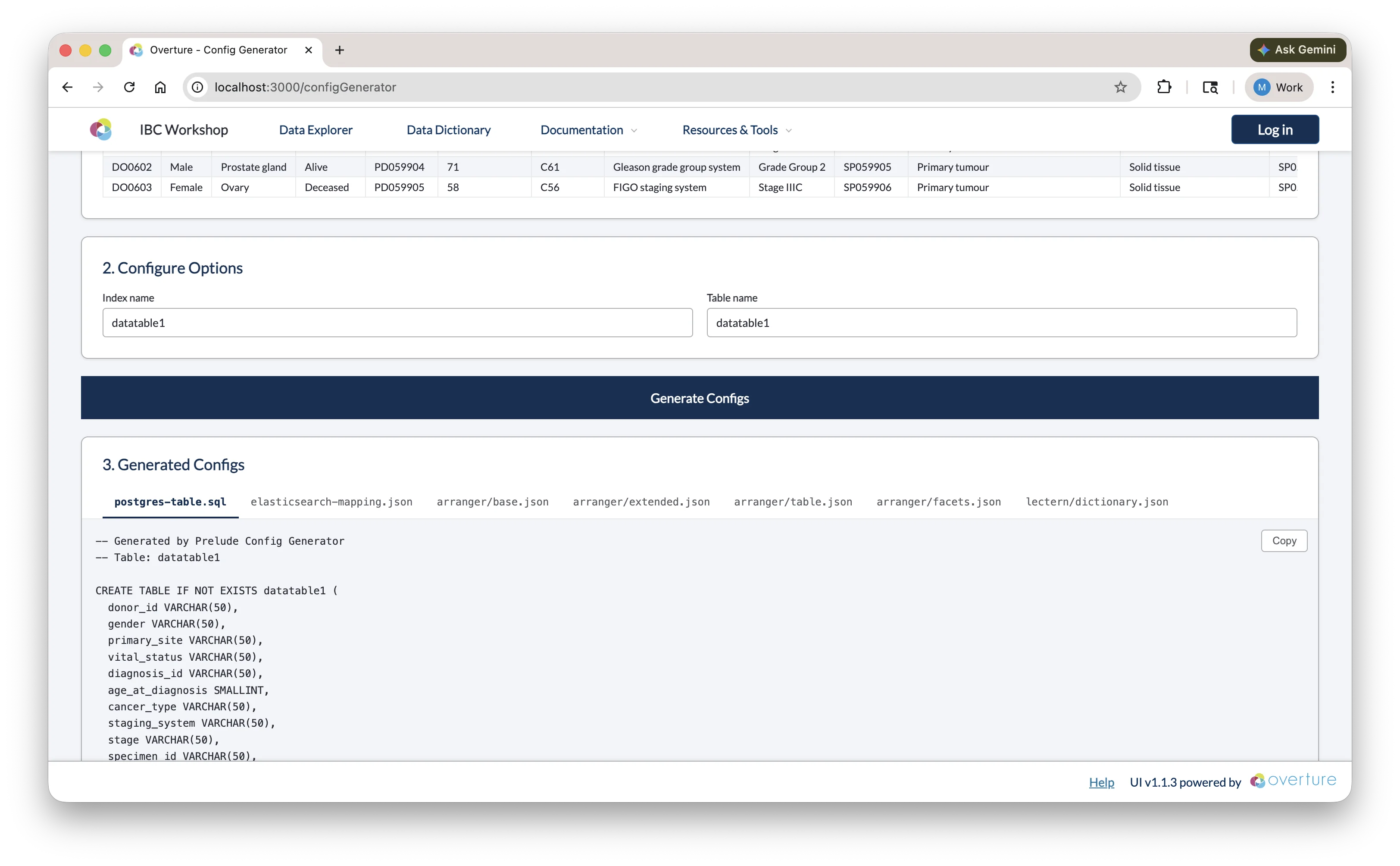
Reviewing the Output
The generated configs are a starting point; review each file before saving and adjust as needed:
postgres-table.sql
The generator infers SQL column types from your data (e.g. VARCHAR for text, SMALLINT or INTEGER for numbers) and adds a submission_metadata JSONB column for tracking. You may want to:
- Increase
VARCHARlengths for columns with longer values - Change
SMALLINTtoINTEGERfor larger numeric ranges - Add constraints if needed (e.g.
NOT NULL)
Click here to see a breakdown of the generated SQL
-- PostgreSQL table creation script
-- Generated from CSV analysis
-- Table: datatable1
-- Columns: 24 (23 data + 1 submission_metadata)
CREATE TABLE IF NOT EXISTS datatable1 (
donor_id VARCHAR(50),
gender VARCHAR(50),
primary_site VARCHAR(50),
vital_status VARCHAR(50),
diagnosis_id VARCHAR(50),
age_at_diagnosis SMALLINT,
cancer_type VARCHAR(50),
staging_system VARCHAR(50),
stage VARCHAR(50),
specimen_id VARCHAR(50),
specimen_type VARCHAR(63),
tissue_source VARCHAR(50),
sample_id VARCHAR(50),
sample_type VARCHAR(50),
treatment_id VARCHAR(50),
treatment_type VARCHAR(50),
treatment_start SMALLINT,
treatment_duration SMALLINT,
treatment_response VARCHAR(50),
drug_name VARCHAR(50),
followup_id VARCHAR(50),
followup_interval SMALLINT,
disease_status VARCHAR(50),
submission_metadata JSONB
);
-- Unique index on submission_id within the JSONB metadata column.
-- Required by Conductor's ON CONFLICT deduplication: if a row with the same
-- submission_id is uploaded again, it is silently skipped instead of duplicated.
DROP INDEX IF EXISTS idx_datatable1_submission_id;
CREATE UNIQUE INDEX IF NOT EXISTS idx_datatable1_submission_id
ON datatable1 ((submission_metadata->>'submission_id'));
| Element | What it means |
|---|---|
VARCHAR(50) | A variable-length text field with a max of 50 characters. Increase this if any values in your column are longer, otherwise PostgreSQL will truncate or reject them on insert. |
SMALLINT | A 2-byte integer supporting values from −32,768 to 32,767. Change to INTEGER (up to ~2.1 billion) or BIGINT if your values can exceed that range. |
submission_metadata JSONB | A binary JSON column added by the generator and populated by Conductor with each record's tracking info: submission_id, source_file_hash, and processed_at. Do not remove; Conductor depends on it to ensure data is never duplicated. |
CREATE TABLE IF NOT EXISTS | PostgreSQL skips creation if the table already exists, so it's safe to re-run the script without accidentally dropping data. |
Unique index on submission_id | Enforces that each uploaded record has a unique submission ID. If Conductor encounters a duplicate during re-upload, it silently skips the row instead of inserting a duplicate or throwing an error. |
The setup service automatically discovers and executes all SQL files in setup/configs/postgresConfigs/ at startup. Ensure your generated SQL file is saved to this directory before launching the platform.
For a full reference on PostgreSQL data types and CREATE TABLE syntax, see the PostgreSQL documentation.
elasticsearch-mapping.json
Field types are inferred from your CSV data. Review and correct:
- Ensure numeric fields are typed as
integerorfloat, notkeyword - Ensure date fields are typed as
date - Categorical fields should remain
keyword
Click here to see a breakdown of the generated mapping
{
"index_patterns": ["datatable1-*"],
"aliases": {
"datatable1_centric": {}
},
"mappings": {
"properties": {
"data": {
"type": "object",
"properties": {
"donor_id": { "type": "keyword" },
"gender": { "type": "keyword" },
"primary_site": { "type": "keyword" },
"vital_status": { "type": "keyword" },
"diagnosis_id": { "type": "keyword" },
"age_at_diagnosis": { "type": "integer" },
"cancer_type": { "type": "keyword" },
"staging_system": { "type": "keyword" },
"stage": { "type": "keyword" },
"specimen_id": { "type": "keyword" },
"specimen_type": { "type": "keyword" },
"tissue_source": { "type": "keyword" },
"sample_id": { "type": "keyword" },
"sample_type": { "type": "keyword" },
"treatment_id": { "type": "keyword" },
"treatment_type": { "type": "keyword" },
"treatment_start": { "type": "integer" },
"treatment_duration": { "type": "integer" },
"treatment_response": { "type": "keyword" },
"drug_name": { "type": "keyword" },
"followup_id": { "type": "keyword" },
"followup_interval": { "type": "integer" },
"disease_status": { "type": "keyword" }
}
},
"submission_metadata": {
"type": "object",
"properties": {
"submission_id": { "type": "keyword", "null_value": "No Data" },
"source_file_hash": { "type": "keyword", "null_value": "No Data" },
"processed_at": { "type": "date" }
}
}
}
},
"settings": {
"number_of_shards": 1,
"number_of_replicas": 0
}
}
| Element | What it means |
|---|---|
index_patterns: ["datatable1-*"] | This template applies to any index whose name starts with datatable1-. Elasticsearch uses it automatically when a matching index is created. |
aliases: { datatable1_centric } | A stable reference name that Arranger queries. Even if you version your indices, the alias always points to the right one; do not change this without also updating base.json. |
data object | All your CSV columns are nested here. This separation keeps your data fields distinct from system metadata. |
keyword type | Exact-match text field, used for categorical values and faceted filtering. Cannot do range queries. |
integer type | Whole number field, supports range queries and numeric aggregations. |
submission_metadata object | Added by the generator, populated by Conductor. Contains submission_id, source_file_hash, and processed_at; do not remove. |
null_value: "No Data" | What Elasticsearch displays when a field has no value. Ensures missing data is visible in facets rather than silently absent. |
number_of_shards: 1 | Number of primary partitions. One is appropriate for local development; increase for multi-node production clusters. |
number_of_replicas: 0 | Number of copies of each shard. Zero is fine for local dev; set to 1+ in production for redundancy. |
Date fields can be problematic. Elasticsearch is strict about date formats; if your data contains dates in mixed formats or includes timezone offsets, indexing will fail. It's safest to normalise all date values to ISO 8601 format (YYYY-MM-DD or YYYY-MM-DDTHH:mm:ssZ). If you're unsure, leaving a date field as keyword will allow it to index without errors, though you'll lose date-range filtering.
For a full reference on Elasticsearch mapping types and settings, see the Elasticsearch mapping documentation.
arranger/base.json
Click here to see a breakdown of base.json
{
"documentType": "records",
"esIndex": "datatable1-index"
}
| Field | What it means |
|---|---|
documentType | Always set to "records" by the generator. |
esIndex | The Elasticsearch index or alias that Arranger queries. Must match the alias defined in your mapping (datatable1_centric). |
Update esIndex to match the alias from your Elasticsearch mapping (datatable1_centric), not the index name pattern (datatable1-*). Arranger queries through the alias so it always points to the correct index.
arranger/extended.json
Display names are auto-generated by converting snake_case to Title Case. Review and adjust any that don't read well; these are the labels shown to users in the portal UI.
Click here to see a breakdown of extended.json
{
"extended": [
{
"displayName": "Donor Id",
"fieldName": "data.donor_id"
},
{
"displayName": "Age At Diagnosis",
"fieldName": "data.age_at_diagnosis"
},
{
"displayName": "Cancer Type",
"fieldName": "data.cancer_type"
}
]
}
| Field | What it means |
|---|---|
fieldName | The full dot-notation path to the field in Elasticsearch (e.g. data.donor_id). Must match the mapping exactly. |
displayName | The label shown to users in the portal UI for this field. Edit these freely; they have no effect on the underlying data. |
For a full reference on extended.json, see the Arranger extended configuration docs.
arranger/table.json
Consider hiding metadata columns (e.g. submission_metadata.*) by setting "show": false and "canChangeShow": false.
Click here to see a breakdown of table.json
{
"table": {
"rowIdFieldName": "submission_metadata.submission_id",
"columns": [
{
"canChangeShow": true,
"fieldName": "data.donor_id",
"show": true,
"sortable": true,
"jsonPath": "$.data.donor_id",
"query": "data { donor_id }"
},
{
"canChangeShow": true,
"fieldName": "data.cancer_type",
"show": true,
"sortable": true,
"jsonPath": "$.data.cancer_type",
"query": "data { cancer_type }"
}
]
}
}
| Field | What it means |
|---|---|
rowIdFieldName | The field used as a unique row identifier. Defaults to submission_metadata.submission_id; do not change. |
fieldName | Dot-notation path to the field in Elasticsearch. Must match the mapping. |
show | Whether the column is visible in the table by default. |
canChangeShow | Whether users can toggle this column's visibility using the column selector. |
sortable | Whether clicking the column header sorts the table. Disable for high-cardinality text fields. |
jsonPath | JSONPath expression used to extract the value from the response. Matches the field structure. |
query | The GraphQL sub-selection used to fetch this field. Must reflect the nested structure in your mapping. |
For a full reference on table.json, see the Arranger table configuration docs.
arranger/facets.json
Remove or deactivate facets that aren't useful for filtering, such as unique ID fields. Reorder entries to put the most important filters at the top.
Click here to see a breakdown of facets.json
{
"facets": {
"aggregations": [
{
"active": true,
"fieldName": "data__gender",
"show": true
},
{
"active": true,
"fieldName": "data__cancer_type",
"show": true
},
{
"active": false,
"fieldName": "data__donor_id",
"show": false
}
]
}
}
| Field | What it means |
|---|---|
fieldName | Double-underscore notation for the field path (e.g. data__cancer_type for data.cancer_type). This is specific to facets.json and differs from the dot-notation used in other config files. |
active | Whether the facet is enabled and will appear in the sidebar. |
show | Whether the facet is visible by default (can be used alongside active to pre-collapse a facet). |
For a full reference on facets.json, see the Arranger facet configuration docs.
Checkpoint
Before proceeding, confirm:
setup/configs/postgresConfigs/datatable1.sqlexists and contains a CREATE TABLE statement matching your CSV columnssetup/configs/elasticsearchConfigs/datatable1-mapping.jsonexists and contains field mappingssetup/configs/arrangerConfigs/datatable1/containsbase.json,extended.json,table.json, andfacets.jsonbase.jsonhas"esIndex": "datatable1_centric"(the alias, not the index name)- You've made at least one change to
facets.json(hidden a facet or reordered entries) setup/configs/lecternDictionary/datatable1.jsonexists (if using the Lectern data dictionary)
Next: We will learn how docker-compose.yml picks up these configuration files and wires them into the running services.